Abstract
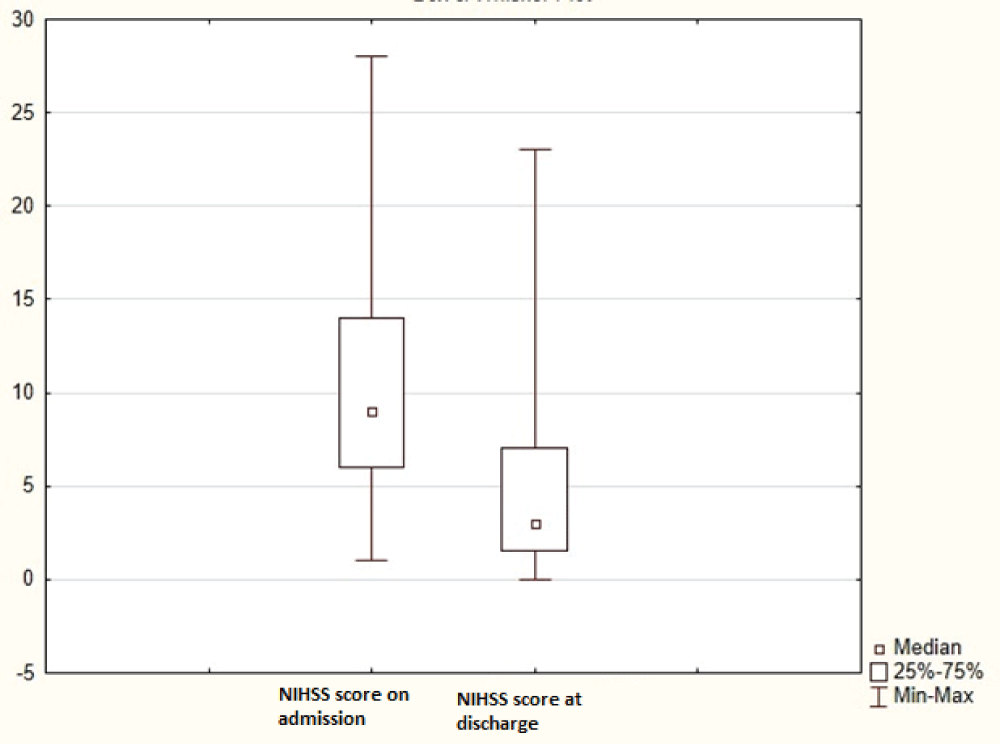
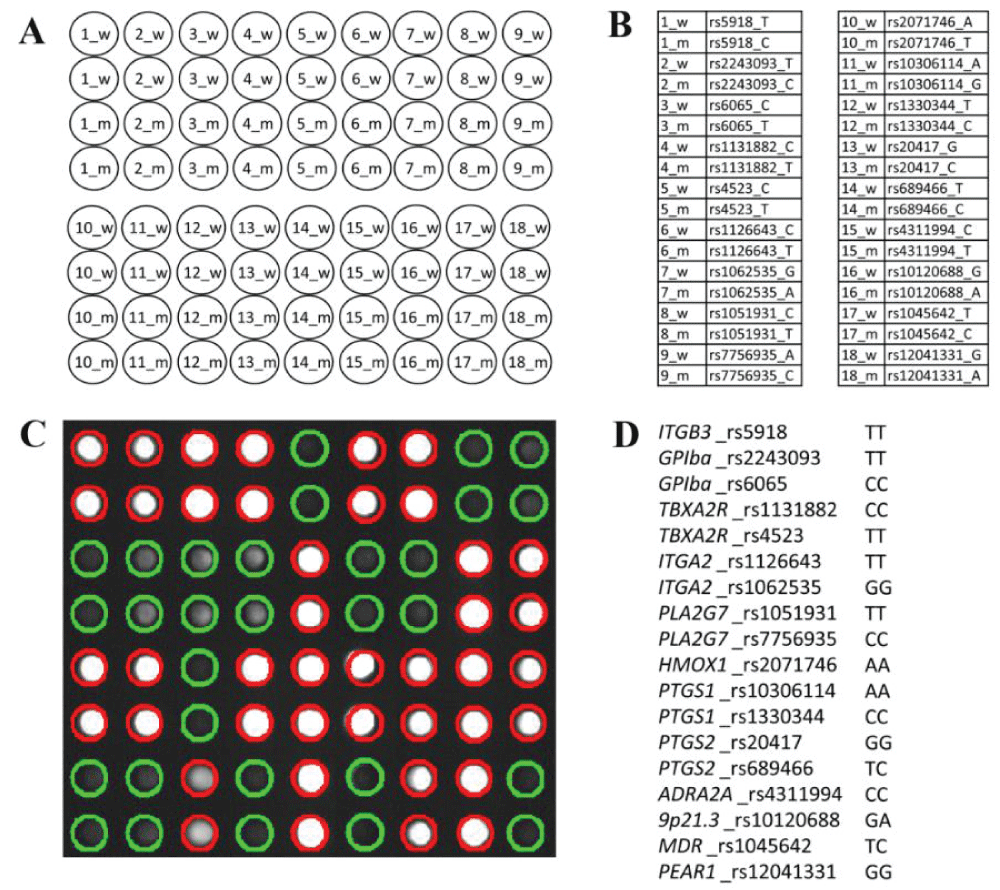
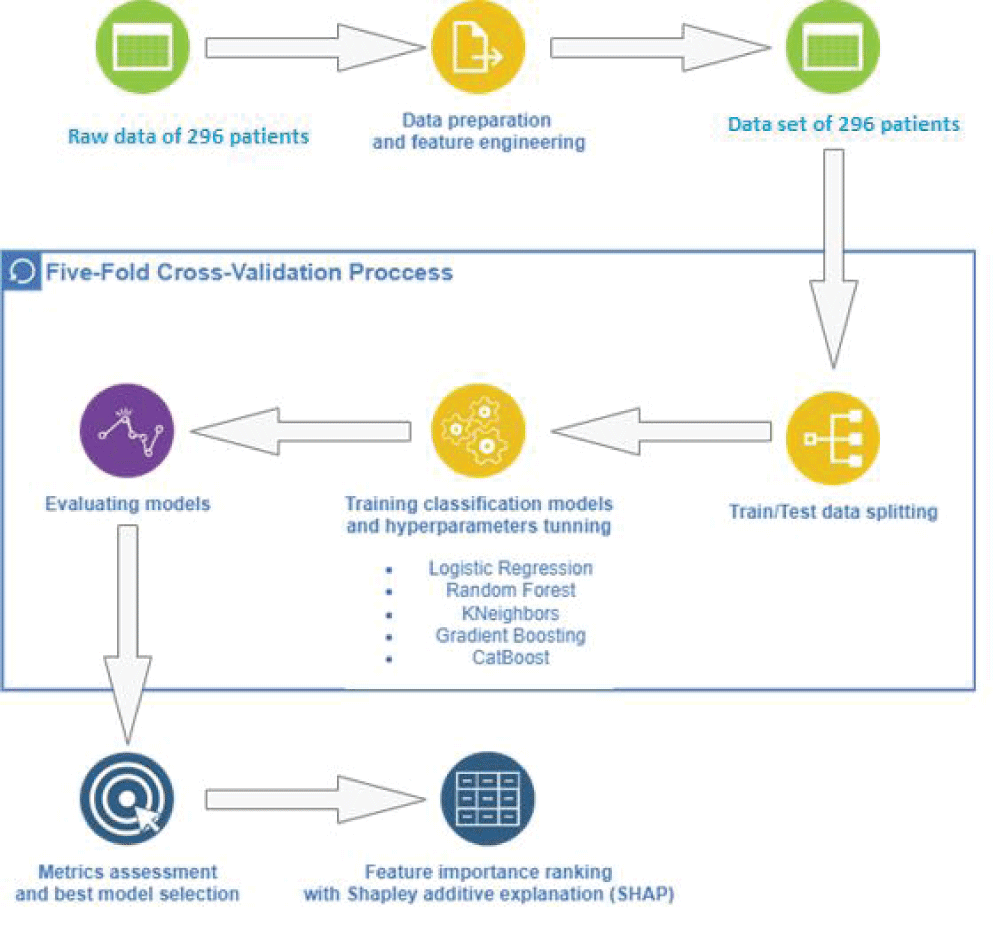
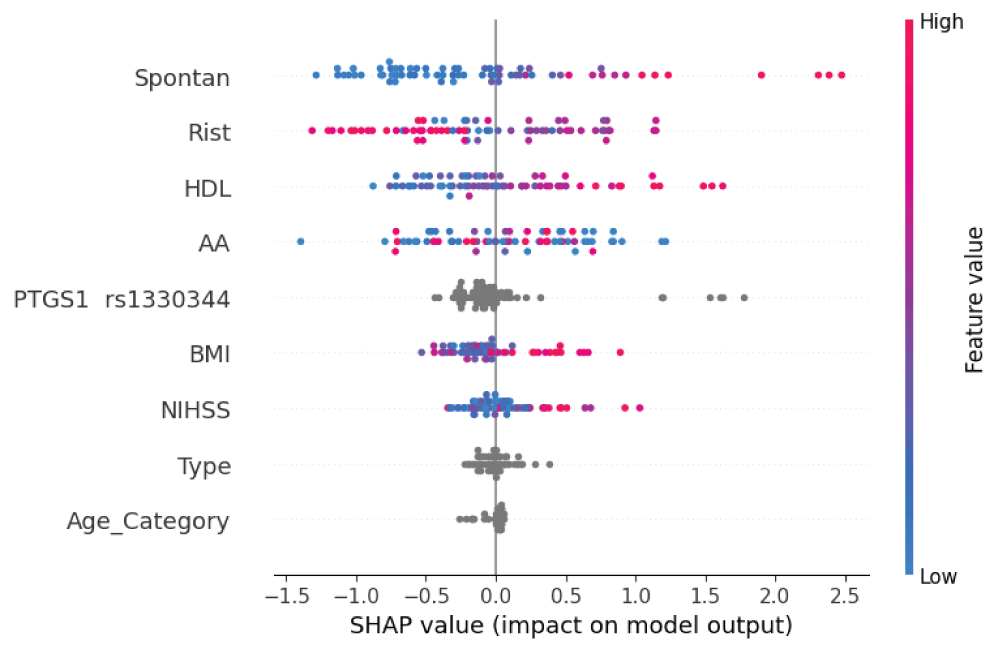
The aim of the study was to determine the incidence of laboratory aspirin resistance; and to study the associations of genetic markers and clinical and laboratory parameters (including parameters of the platelet hemostasis) in patients with non-cardioembolic ischemic stroke using machine learning methods to assess the prognosis of recurrent ischemic strokes. Clinical and laboratory data (including induced platelet aggregation) were analyzed from 296 patients with ischemic stroke who were treated in the stroke center of City Clinical Hospital No. 1 named after. N.I. Pirogov. The frequencies of polymorphic variants of the ITGB3, GPIba, TBXA2R, ITGA2, PLA2G7, HMOX1, PTGS1, PTGS2, ADRA2A, ABCB1, PEAR1 genes and intergenic region 9p21.3) in patients with non-cardioembolic ischemic stroke, which were identified using hydrogel biochip technology, were determined. Using the developed machine learning model, additional clinical and genetic factors influencing the development of laboratory aspirin resistance and recurrent ischemic stroke were studied. In the future, the identified factors can be used for differentiated prevention of recurrent ischemic strokes.